By Molnár-Institute for…
MOLNÁR-INSTITUTE DryLab®4 LC modelling helps in scouting for cytotoxic metabolites from Brazilian sponge
The Design of Experiments (DoE) and multi column screening capabilities of Molnár-Institute’s proprietary software suite, DryLab®4, have proved highly useful in developing a faster and more efficient liquid chromatography workflow for identifying complex networks of metabolic cytotoxins in organic substances, using a Brazilian marine sponge as a study.
The research paper ‘Integrated analytical workflow for chromatographic profiling and metabolite annotation of a cytotoxic Phorbas amaranthus extract’; is published in the Journal of Chromatography [1].
DoE capabilities
The study, led by Dr. Bruno do Amaral and Prof. Dr. Quezia Cass with scientists from four Brazilian academic institutions; Department of Chemistry, Federal University of São Carlos, Natural Products Research Institute, Federal University (UFF) Rio de Janeiro, Department of Marine Sciences, Institute of the Sea, UFF São Paulo, and the Natural Products Laboratory, Institute of Chemistry, UFF Niteroi [1].
They set out to achieve fast and efficient chromatographic separations to identify metabolites present in Phorbas amaranthus, a genus of marine sponge found in Brazilian waters and known to be a rich source of bioactive cytotoxins. To make this possible, the team planned to use the Design of Experiments (DoE) capabilities of the DryLab® platform and a dedicated UHPLC control software, to create a usable scouting liquid chromatographic system that would eliminate redundant experiments.
Faster separation
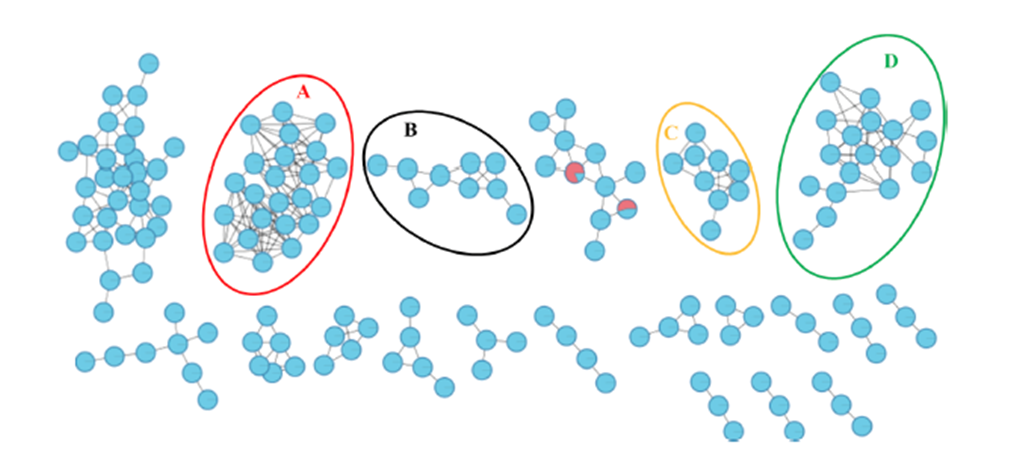
The team started from a crudely prepared and partitioned hydroalcoholic extract of the sponge Phorbas amaranthus collected from the Brazilian coast. They used tumor cell culture models to evaluate cytotoxicity of the crude extract and its fractions. Fragmentation pathways assembled from HRMS data allowed them to annotate 18 known Phorbas metabolites, and infer 17 more, based on Global Natural Product Social Molecular Networking (GNPS), matching with a further 29 metabolites annotated through a molecular subnetwork.
The main benefit of perfecting DoE with the aid of DryLab® software was to obtain faster and more efficient chromatographic separation of the active hexane fraction, further enabling untargeted high-resolution mass spectrometry (HRMS) data.
Finding new metabolites
The crude extract obtained from the marine sponge P. amaranthus displayed cytotoxicity against the tumor cell line HCT 116 and a partition in hexane (PaHex) revealed comparable bioactivity to that of the crude extract. The automation in column and eluent scouting associated with DryLab® optimization meant that only 12 experiments (24 chromatographic runs) were required to produce a metabolite profile with selectivity and short analysis time that enabled the dereplication by LC-HRMS of 64 annotated metabolites from the PaHex, including eight identified for the first time and ten not hitherto associated with sponges.
The team succeeded in developing a chromatographic method workflow, accelerated by the use of automated scouting systems for selecting the DoE parameters and DryLab® for predicting the optimized chromatographic conditions. This has proved useful for profiling natural product libraries, as well as data curation by molecular clusters to form a valuable new tool for natural product chemists.
“Besides the analytical workflow proposed herein, this study has also contributed to the metabolic profiling of P. amaranthus and can be employed to guide the isolation of cytotoxic metabolites in complex natural extracts,” the study concludes.
Reference
[1] Do Amaral, B.S., Da Silva, F.B., Leme, G.M., Schmitz, L.S., Jimenez, P.C., Campos Martins, R.C., Cass, Q.B., Valverde, A.L., (2021) ‘Integrated analytical workflow for chromatographic profiling and metabolite annotation of a cytotoxic Phorbas amaranthus extract’; Journal of Chromatography B, Vol. 1174, 122720. DOI: https://doi.org/10.1016/j.jchromb.2021.122720
Resources
Click on Integrated workflow for chromatographic profiling of Phorbas amaranthus metabolites to download full study.