By ATG:biosynthetics GmbH
PepID for Epitope mapping
Introduction
ATG:biosynthetics has developed its PepID technology to provide advantageous, straightforward workflows for epitope analyses of antigens and antigenic determinants.
PepID’s strong computational algorithms generate rationally designed, deterministic DNA-encoded peptide libraries. The system hosts and expresses these libraries using a smart bio-based vector system, generally based on proteins that come from diagnostically and epidemiologically relevant bacteria and viruses, such as flaviviruses, or even cancer antigens.
In silico, in vitro, in vivo
PepID is a bio-peptide system in which peptides are produced and surface displayed in a biological system, usually a bacterial host organism or Mammalian cells, from a plasmid DNA construct. It streamlines the identification of epitopes as well as the development of bioanalytics, diagnostics and vaccines using in vitro chemical peptides and in vivo bio-peptides.
To identify B-cell, linear or conformational, or T-cell epitopes, ATG performs global data mining and analyses the available literature data and sequences of the proteins of interest to establish optimized non-redundant sets of peptides mirroring the biodiversity of relevant potential antigens.
In a two steps procedure, the analysis system uses designs of chemical peptide microarrays covering the pool of synthetic peptides previously determined in silico to identify all potential epitopes in an in vitro pre-screening. Following this first evaluation of the level of recognition of each peptide by the serum of interest, PepID gathers desired candidates in library for further display as bio-peptides for in vivo epitope-mapping which mimic the natural context of the epitope in the cellular environment.
Epitope mapping deterministic library
PepID thus generates libraries that are deterministic as the peptide sequences are based on existing and usually well-studied proteins. Compared with chemically synthesized random libraries, PepID provides permutations of sequences that are sorted with far more biological relevance.
The PepID library feature delivers numerous advantages:
- Reproduction of unlimited peptide candidates without any alteration.
- High flexibility in expression through the library’s ability to use simple sub cloning to adapt tags or inducibility to fit with the project’s needs.
- Radically reduced library size compared to random libraries with defined low complexity.
- Elimination of irrelevant peptides or empty vectors.
Optimal workflow
PepID combines high-throughput screening potential of in vitro synthetic chemical peptides gathered in microarray with the flexibility and stringent reproducibility of in vivo bio-eptides library design. Together, these define optimal workflows for identification of new epitopes and the development of bioanalytics, diagnostics and vaccines.
Comparative genomics is an indispensible tool in project planning, fitting the functional realities of specific activities with protein expressing systems.
PepID technology can be used to perform mapping of epitopes, covering the biodiversity of pathogen bacterial and viral genomes in order to discover the most plausible antigens.
If you wish to cover specific immunologically more relevant areas more tightly, you can adjust either the peptide length or the peptide overlap as shown in Figure 2. This can help you identify new linear epitopes and possibly even conformational epitopes.
Resources
Click on PepID for Epitope mapping for more information.
Click on ATG to contact the company directly.
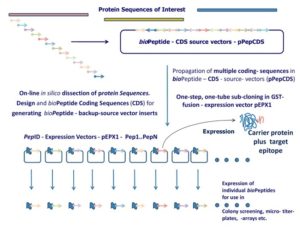
Figure 1: PepID separates hosting and expression into two subsystems. First, a protein sequence of interest is reverse translated and then partitioned into linear peptides of defined length and overlap in silico. Peptide-coding DNA sequences are then assembled into a single or a set of compact constructs to be synthesized de novo and hosted on the source / back-up vector from which they can be released whenever necessary.
Peptides are released by simply cutting with a unique combination of restriction enzymes and then cloned into the expression vector. The expression vector can be uniquely tagged (e.g. GST) to produce a fusion peptide ready for expression testing and utilization.
Supplier Information
Supplier: ATG:biosynthetics GmbH
Address: Weberstrasse 40, 79249 Merzhausen, Germany
Tel: +49 (0) 761 888 9424
Fax: +49 761 888 9425
Website: www.atg-biosynthetics.com/