By Bachem AG
Diagnostic peptides
Use of peptides as diagnostic tools is evolving to enable simpler, more accessible and faster testing procedures for both biomarkers and epitopes. This in turn is allowing fast and accurate diagnosis of diseases beyond laboratories and well-equipped healthcare facilities. Current diagnostic peptide clinical developments are focusing on point-of-care diagnosis, which has several benefits:
- Higher chance of fighting the disease at early stage and therefore shortening its duration.
- Reduced risk of trial-and-error treatment and over-prescription.
- Scope for more specialized therapy (e.g. avoiding broad-spectrum antibiotics).
- Faster quarantining of infectious patients, lowering risk of nosocomial infections by faster diagnosis. (Estimates suggest the Sierra Leone Ebola epidemic of 2014 could have been reduced by up to 30% in scale with rapid testing methods).
Recent research has developed novel materials to improve disease diagnosis through the detection of peptide biomarkers. Peptide sequences are part of larger proteins, which constitute the most abundant and versatile biomolecules present in living cells, with properties dictated by genetic code. Their diverse structure and function are responsible for key cellular processes including metabolism and cell movement. They are also implicated in many diseases whether genetic, multifactorial or degenerative. Studying protein function and shape is central for understanding disease and for the discovery of new drugs.
Use of peptides in diagnosis
Peptides play an important role in the detection of diseases and can be grouped into two categories based on the role they play in the diagnosis.
Biomarkers: Proteins that contain biomarkers exhibit changes in concentration and/or degradation in body fluids change that indicate disease. Bachem offers a range of such biomarker peptides (see Resources). The determination of these concentrations usually requires a well-equipped laboratory and the transport of the samples, which is detrimental to a fast diagnosis.
Epitopes: Peptides that are epitopes of pathogens offer potential for rapid diagnosis by detecting antibodies that have been produced by the human body as a response to the antigens.
The standard method for detecting antibodies using diagnostic peptides is an enzyme-linked immunosorbent assay (ELISA). Although the exact method can differ from case to case, the general procedure is as follows:
- Link peptide sequences specific to antibody of interest to the walls of a microplate, in which the residual binding sites are then blocked.
- Wash microplate with the analyte, causing present antibodies to bind to the bound peptides.
- Rinse microplate with mild detergent, removing any unbound biomolecules.
- Test microplate for the presence of bound antibodies (e.g. using an anti-antibody tagged with a fluorophore), which would indicate the presence of a disease.
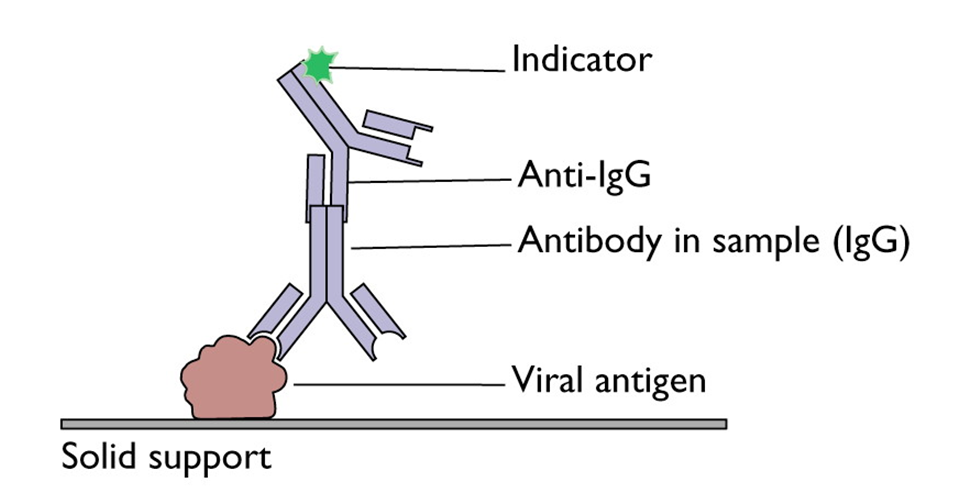
The final state of a successful antibody detection is summarized in Figure 1.
Usually the plates can afterwards be washed with a washing solution, which removes the antibodies (e.g. containing 2-mercaptoethanol), allowing the same plate to be used several times.
Modern peptide diagnosis
The above methodology has been established since the 1970s. However, developments over the past two decades have enabled much faster discovery of new potentially useful epitopes, via:
- Structural elucidation of complete pathogenic proteins, made possible by advances in proteomics
- High throughput screening of standardized microwell plates, providing a rapid mechanism for immunological testing of epitopes
The large amount of epitope data now available require fast and efficient approaches to find the best candidates.
One way to determine which part of a pathogen sequence binds to a specific antibody is to test a set of short (10-18 AAs) overlapping peptide sequences for binding to the antibody. This method allows the detection of linear antibody binding epitopes. For complex pathogens, this of course requires a very large library of synthetic peptides, which is why in these cases bioinformatics tools are used to determine epitope regions of interest.
The use of short peptide sequences offers the advantage of easier synthetic access but does unfortunately come with drawbacks. Many epitopes are either non-linear or influenced by the three-dimensional confirmation of the peptide chain. In this case, it might be possible to detect binding epitopes using longer peptides (up to 30 AAs) for the screening or again by using bioinformatics tools.
The challenge in designing diagnostic peptides is to find sequences that not only bind strongly to the antibody of interest (high sensitivity), but also only to this antibody (high selectivity). Binding to other antibodies will lead to cross-reactivity, which in turn leads to false positive results and potentially treatment of the wrong disease. Another example of problematic interference is the detection of tuberculosis (TB) antibodies in patients who are also HIV positive. Due to the alteration of the normal host immune response in persons infected with HIV, antibody tests for TB can return false negative results, since the B-cell relevant B-cell epitopes differ in comparison to HIV negative patients. This leads to a potentially deadly delay in treatment and the risk of infection of other patients. The case of TB in conjunction with HIV is especially problematic since in low-income countries both diseases are prevalent.
It is therefore necessary to check candidate sequences against such cross-reactivity to ensure the accuracy of the test even in case of multiple simultaneous diseases. This is another area where bioinformatics plays an increasingly important role. With a given sequence and epitope databases such as the IEDB algorithms can predict antibodies with the potential of cross-reactivity with this sequence, which can then be tested by an ELISA.
Despite all these obstacles, antibody detection has emerged as an important diagnostic tool. Examples of successfully established procedures are the rapid plasma reagin test (RPR) for syphilis as well as the rapid HIV-test. Both tests do however require follow-up tests for confirmation in the case of positive results.
Custom peptide synthesis
An important factor for the reproducibility of antibody detection tests is the purity of the peptide that is used. Synthetic peptides are at an advantage in this regard compared to natural or recombinant biomolecules/pathogens, which can also be used to detect antibodies. Synthetic peptides can be produced in a highly reproducible manner, which is crucial for the commercialization of a diagnostic method using peptides. Bachem offers a custom synthesis service, which is ideally suited for our partners in the diagnostic business.
The Bachem Center of Excellence for Custom Synthesis in St. Helens (UK) is certified according to ISO 13485 for the manufacture of peptides as raw materials for medical devices. It offers a state of the art quality control system as well as additional services such as dedicated equipment and change control.
References
S.J.Carmona, P.A.Sartor, M.S.Leguizamón, O.E.Campetella, F.Agüero, Diagnostic peptide discovery: prioritization of pathogen diagnostic markers using multiple features. PLOS ONE 2012, 7, e50748.
R.H.Meloen, W.C.Puijk, J.P.M.Langeveld, J.P.M.Langedijk, P.Timmerman, Design of synthetic peptides for diagnostics. Current Protein & Peptide Science 2003, 4, 253-260.
K.A.Navalkar, S.A.Johnston, P.Stafford, Peptide based diagnostics: Are random-sequence peptides more useful than tiling proteome sequences? Journal of Immunological Methods 2015, 417, 10-21.
P.Nouvellet, T.Garske, H.L.Mills, G.Nedjati-Gilani, W.Hinsley, I.M.Blake et al, The role of rapid diagnostics in managing Ebola epidemics. Nature 2015, 528, S109.
G.Shen, D.Behera, M.Bhalla, A.Nadas, S.Laal, Peptide-based antibody detection for tuberculosis diagnosis. Clinical and Vaccine Immunology 2009, 16, 49-54.
R.E.Soria-Guerra, R.Nieto-Gomez, D.O.Govea-Alonso, S.Rosales-Mendoza, An overview of bioinformatics tools for epitope prediction: Implications on vaccine development. Journal of Biomedical Informatics 2015, 53, 405-414.
Resources
Click on Bachem peptide biomarkers to download brochure.
Click on Bachems News to see latest News & Events.